Exploring genomes to understand genetic instability
|
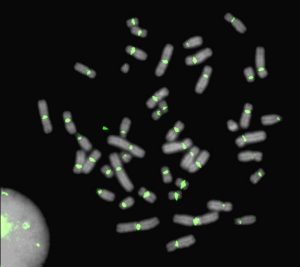
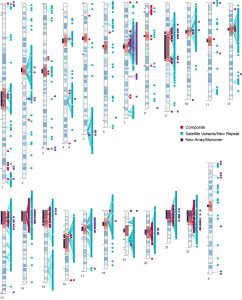
The development of genome-scale sequence data has provided a foundation for studying biological processes in targeted species, while the ability to compare the genomes of different species has afforded a greater understanding of genome evolution and the functional elements dispersed among coding and non-coding regions. We are capitalizing on the advantages in employing comparative genomics approaches to understanding genome and species biology to understand lability in centromeres, telomeres and chromosome breakpoints. Within marsupials, the macropodids (kangaroos and wallabies) exhibit rapid chromosome evolution while a sister group, the dasyurids, are chromosomal stable. We are using comparative genome assemblies to build genetic and epigenetic models to uncover the genomic features that guide stability vs instability. How do specific repeats and epigenetic features mediate genome stability? |
 |
Revealing genomic mechanisms of rapid chromosomal speciation
 |
Through comparative genomics and cytogenetics, we have brought to bear newly developed genome assembly tools on a non-traditional model system that exhibits rapid karyotypic evolution: the lesser apes - gibbons. We aim to address key questions on the role retroelements play in rapid chromosome change, centromere function and speciation. How does the activity of retroelements impact chromosomal speciation? |
Dissecting centromere establishment and assembly in humans
Using comparative genomics to understand adaptive responses within marine ecosystems
 |
Despite the advantages in employing comparative genomics approaches to understanding genome and species biology, few species that occupy pivotal roles in marine ecosystems and pelagic food webs have been utilized as model organisms for genome-scale studies. Employing diverse technologies for the development of reference genome assemblies, we have targeted several species for comparative genomics that will afford a greater understanding of adaptive responses to environmental variation and the evolution of novel phenotypes within marine ecosystems. One of our target species is the Southern Ocean salp, Salpa thompsoni, which has shown altered distribution and abundance in Antarctic Ocean ecosystems in recent years. With a high-quality genome assembly, we can begin to unravel the genomic features that allow the salp to adapt to our warming oceans with the devastating consequence of depleting the ocean of the extraordinary diversity of organisms that rely on krill. |